
Accurate comparisons of protein binding sites is critical as structural similarities between binding sites have been associated with similar ligand binding mechanisms. SiteContext compares binding sites using a transportation-theoretic measure computed over a rich set of 3D molecular descriptors. To improve the efficiency of the computations, the Sinkhorn algorithm is applied. These descriptors include steric (shape) features and physicochemical properties, such as partial charges, important to binding. The steric features are computed by binning atomic distributions into spherical histograms centered on each binding site atom. Within each histogram, bins store both the atom count and the average vector from the binned atoms to the center atom. The design of the steric descriptor is motivated by our prior work on non-sequential structure alignment [1].
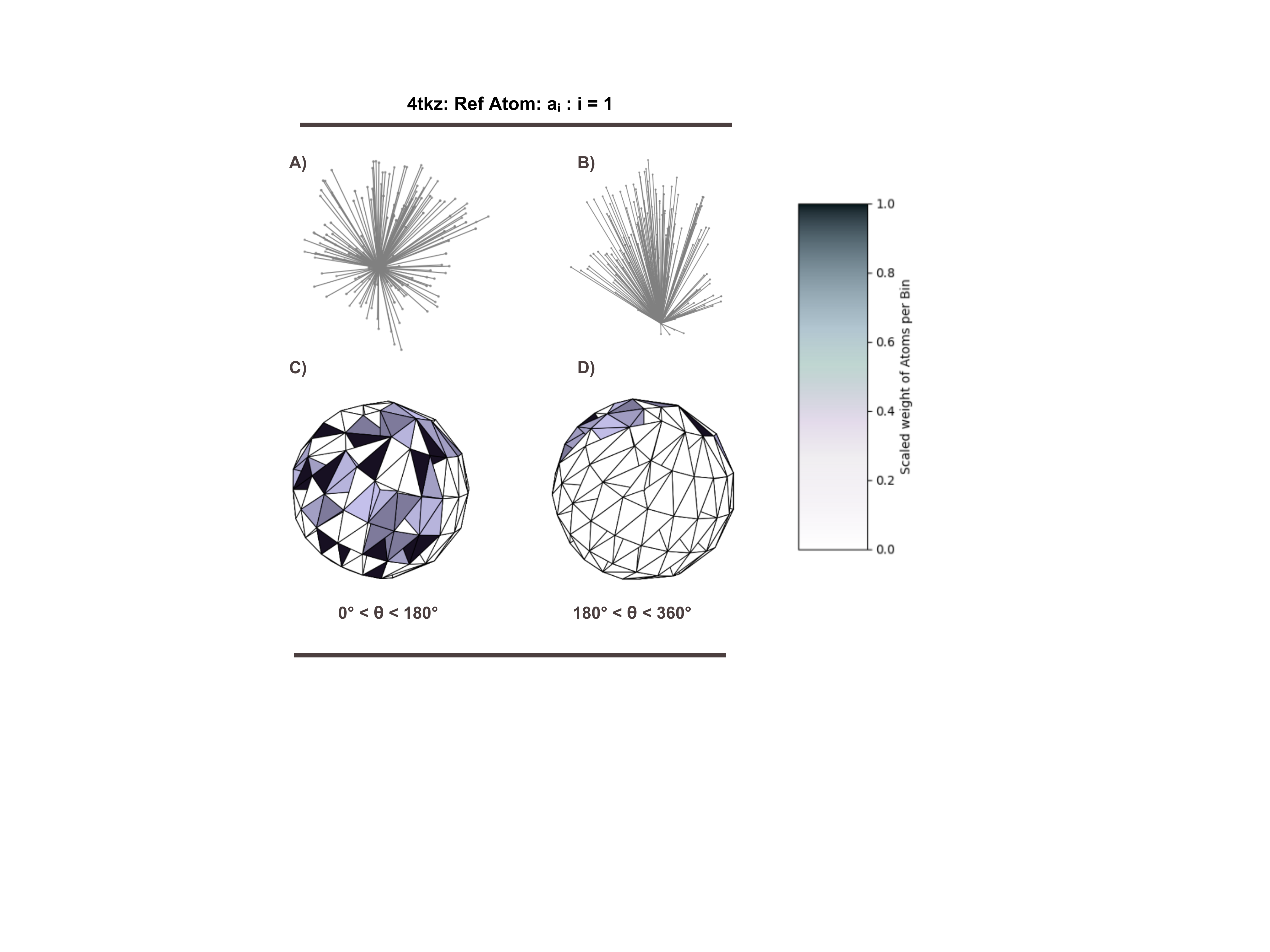
Figure 1: An example of a histogram constructed around reference atom a₁ of the binding site from structure 4ktz is shown. The vectors from the reference atom to all other atoms are illustrated in a) and b), while the corresponding histograms appear in c) and d). The difference between subfigures a) and c) and subfigures b) and d) lies in the range of the azimuthal angle, θ. For better visualization, the weight of each bin was scaled by taking the natural logarithm of non-zero entries and then normalizing the resulting values.
Reference
Please contact Jiadong Yu (Arnold), Hiba Bhatti or Dr. Rahul Singh for any queries regarding this project.
Jiadong Yu (Arnold): jiadong-yu at uiowa dot edu
Hiba Bhatti: hiba.bhatti at gmail dot com
Dr. Rahul Singh: rahul-singh at uiowa dot edu