
Example alignment and their functional inference
The final correspondence examined the protein MAO-A derived from humans (PDB code: 2bxs) and from rats (PDB code: 1o5w), both bound to clorgyline. Despite minor differences in cavity volume, the binding sites share conserved residues. The Sinkhorn-derived EMD score for this pair was 4.75, while the RMSD was 1.63 Å. Both values are relatively low, demonstrating the method's ability to detect structural conservation across species. Further, there were fewer errors in the correspondences, defined as lines being closer together and not matched across the cavity. This occurs because, in highly similar sites, key functional residues are already well aligned, reducing the need to match atoms across the cavity to establish correspondence.
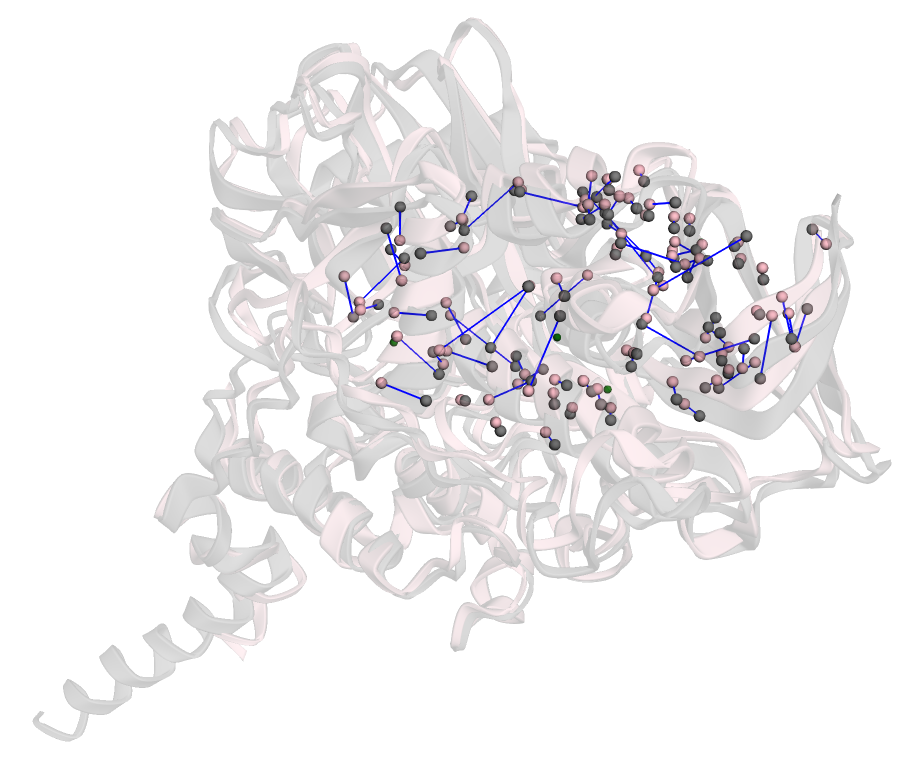
Figure 1: The correspondences between atoms in sites belonging to MAO-A (2bxs) and MAO-A (1o5w).
LSD1 (PDB code: 2z3y), a protein implicated in prostate cancer [1], and human-derived MAO-A (PDB code: 2bxr), associated with Alzheimer's disease, were compared. The two sites share structural similarities but differ in the types of residues within the binding site. This pair was classified as dissimilar, as experimental evidence supports this distinction—LSD1 was shown to bind an inhibitor, whereas MAO-A did not. The proposed method successfully captured subtle differences arising from variations in atomic structure among the differing residues. The overall matching cost and RMSD for this pair were 8.66 and 2.54 Å, respectively—both higher than those of site pairs classified as similar. Notably, there was a high frequency of atoms matched across the cavity, which is less common in similar pairs.
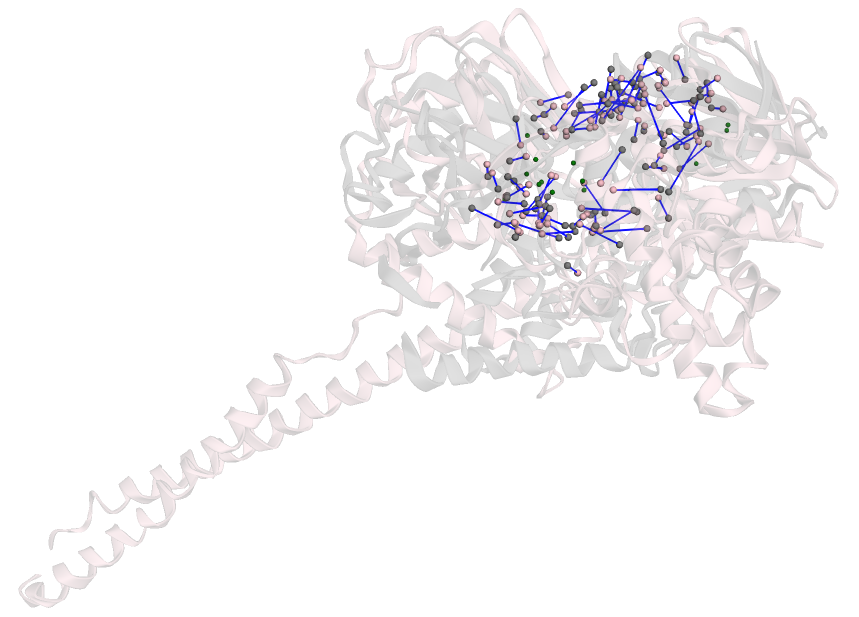
Figure 2: The correspondences between atoms in sites belonging to LSD1 (2z3y) and MAO-A (2bxr).
Reference
Structure Correspondence Viewer